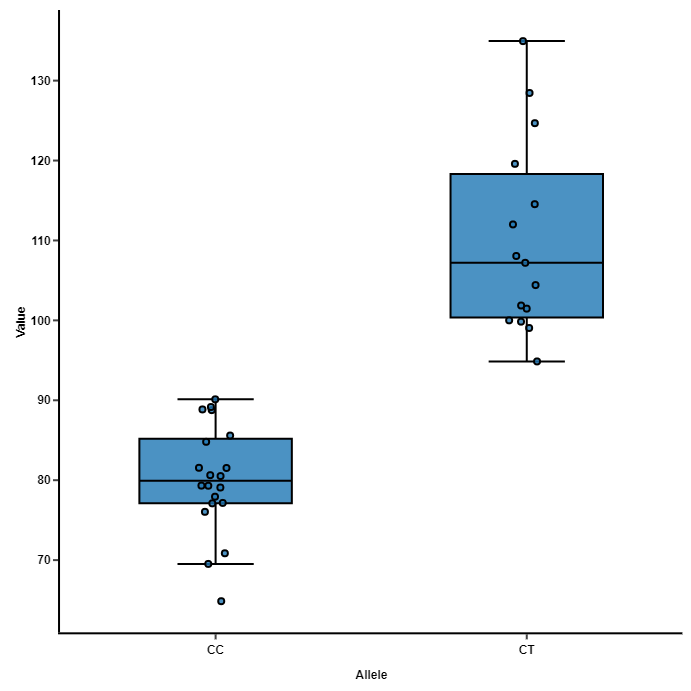
r - How to test overlapping confidence intervals to see a significant difference between groups in a COX propotional model - Cross Validated
![Request] Add elevation reading to Trail Markers. Using Ziplines is a pain when you don't know the elevation. The image is a mock-up of how it could look. : r/GroundedGame Request] Add elevation reading to Trail Markers. Using Ziplines is a pain when you don't know the elevation. The image is a mock-up of how it could look. : r/GroundedGame](https://preview.redd.it/request-add-elevation-reading-to-trail-markers-using-v0-0mrut1cosjx91.png?width=640&crop=smart&auto=webp&s=a476fcaae7fd5dfae58e0b65edbfb340f0a315ed)
Request] Add elevation reading to Trail Markers. Using Ziplines is a pain when you don't know the elevation. The image is a mock-up of how it could look. : r/GroundedGame
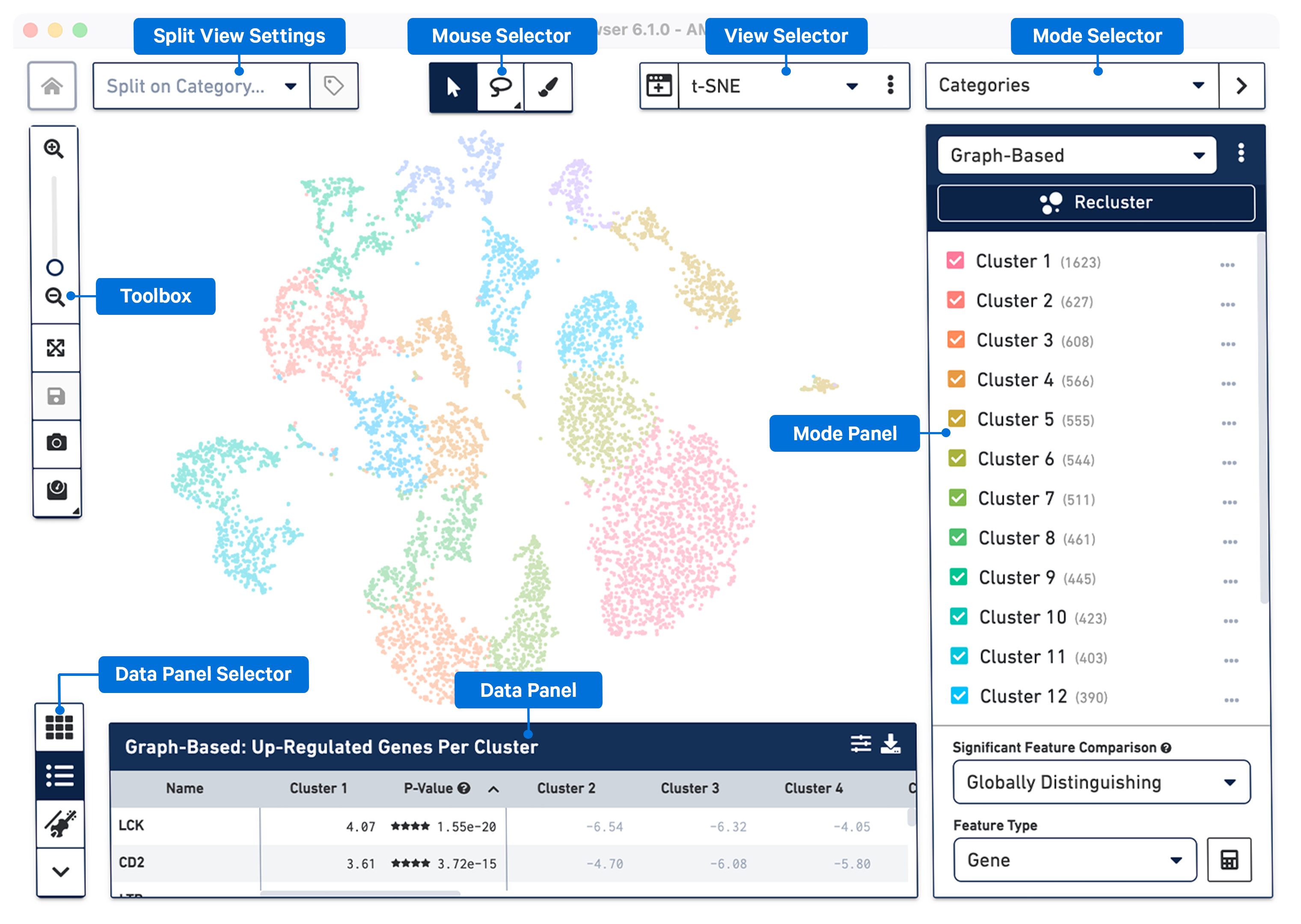
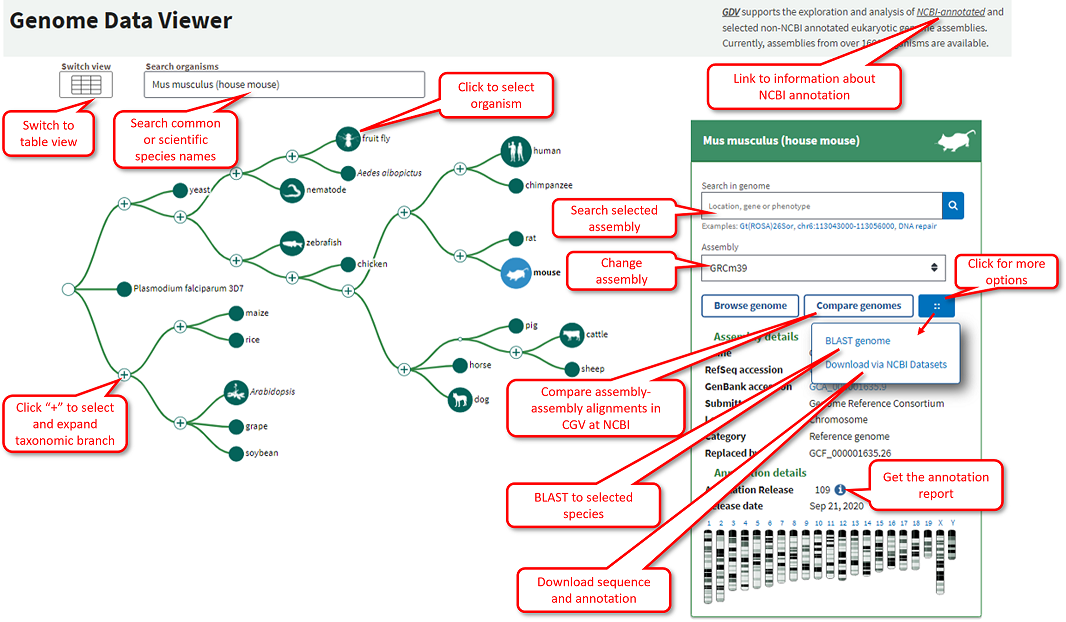
Navigating the Loupe Browser User Interface -Software -Single Cell Gene Expression -Official 10x Genomics Support
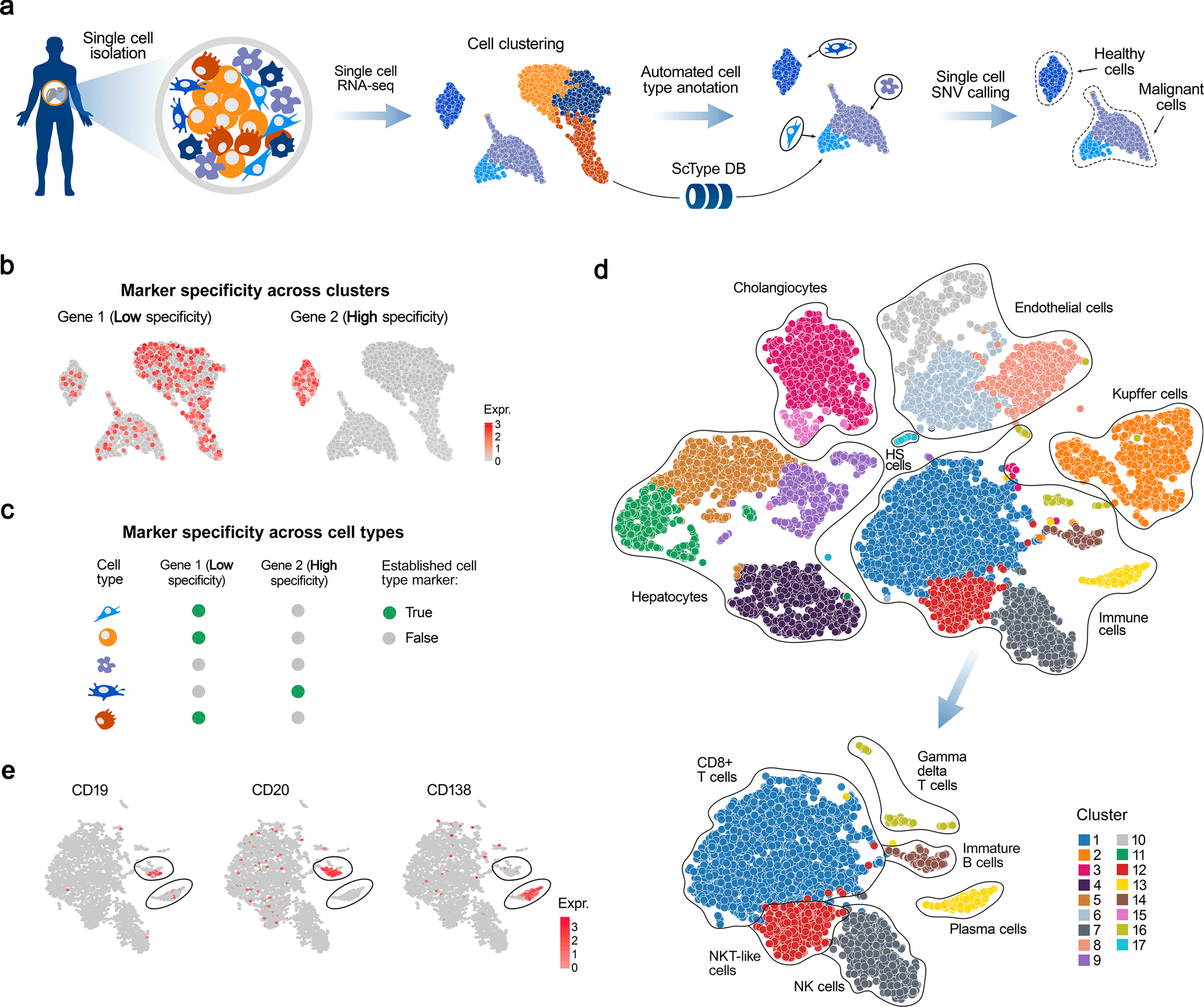
Fully-automated and ultra-fast cell-type identification using specific marker combinations from single-cell transcriptomic data | Nature Communications